scanpy.tl.embedding_density#
- scanpy.tl.embedding_density(adata, basis='umap', *, groupby=None, key_added=None, components=None)[source]#
Calculate the density of cells in an embedding (per condition).
Gaussian kernel density estimation is used to calculate the density of cells in an embedded space. This can be performed per category over a categorical cell annotation. The cell density can be plotted using the
pl.embedding_densityfunction.Note that density values are scaled to be between 0 and 1. Thus, the density value at each cell is only comparable to densities in the same category.
Beware that the KDE estimate used (
scipy.stats.gaussian_kde) becomes unreliable if you don’t have enough cells in a category.This function was written by Sophie Tritschler and implemented into Scanpy by Malte Luecken.
Array type support# Array type
supported
… experimentally in dask
Array✅
❌
❌
❌
- Parameters:
- adata
AnnData The annotated data matrix.
- basis
str(default:'umap') The embedding over which the density will be calculated. This embedded representation is found in
adata.obsm['X_[basis]']`.- groupby
str|None(default:None) Key for categorical observation/cell annotation for which densities are calculated per category.
- key_added
str|None(default:None) Name of the
.obscovariate that will be added with the density estimates.- components
str|Sequence[str] |None(default:None) The embedding dimensions over which the density should be calculated. This is limited to two components.
- adata
- Return type:
- Returns:
Sets the following fields (
key_addeddefaults to[basis]_density_[groupby], where[basis]is one ofumap,diffmap,pca,tsne, ordraw_graph_faand[groupby]denotes the parameter input):adata.obs[key_added]numpy.ndarray(dtypefloat)Embedding density values for each cell.
adata.uns['[key_added]_params']dictA dict with the values for the parameters
covariate(for thegroupbyparameter) andcomponents.
Examples
import scanpy as sc adata = sc.datasets.pbmc68k_reduced() sc.tl.umap(adata) sc.tl.embedding_density(adata, basis='umap', groupby='phase') sc.pl.embedding_density( adata, basis='umap', key='umap_density_phase', group='G1' )
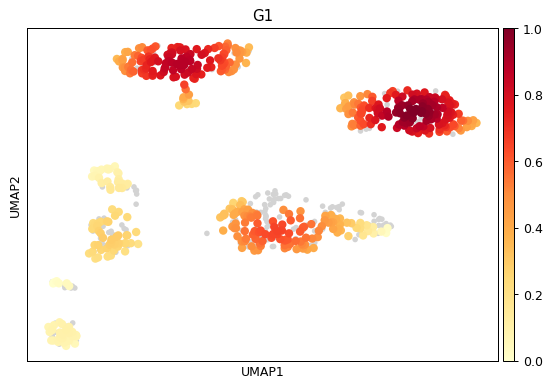
sc.pl.embedding_density( adata, basis='umap', key='umap_density_phase', group='S' )
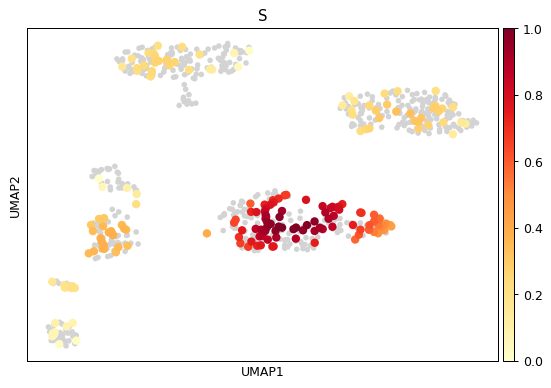
See also